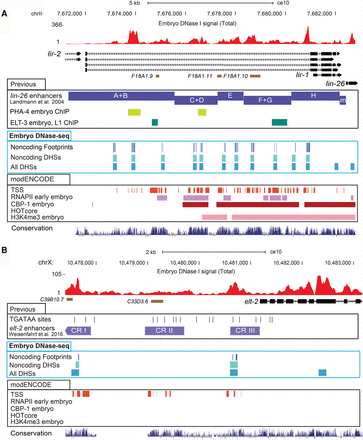
Noncoding DHSs recapitulate many known CRMs. Total DNase signal (red) from both strands of embryo read data shown. Noncoding DHSs (light blue boxes) and all DHSs (medium blue boxes) and TF footprints (dark blue boxes) detected. Additional tracks shown are C. elegans RefSeq genes (black boxes with arrows), noncoding transcripts (brown boxes), and phyloP conservation (very dark blue). Other comparison tracks include TSS (dark orange boxes) (Chen et al. 2013), RNAP II ChIP-seq (red boxes), H3K4me3 (pink), and CBP-1 (lavender boxes) ChIP-chip from modENCODE embryo data (Gerstein et al. 2010). (A) All five known enhancers of lin-26 in the 11-kb first intron of lir-1 are recovered, each harboring at least one embryo noncoding DHS and footprint. Multiple noncoding DHSs are detected upstream of lin-26, in the first intron of lir-1, which harbors known CRMs active in embryos: A+B, C+D, E, F+G, and H (purple boxes) (Landmann et al. 2004). Noncoding DHSs and footprints are detected in each known CRM, and in the case of A+B and C+D, the noncoding DHSs and footprints overlap with PHA-4 ChIP-seq peaks (light green) (Zhong et al. 2010). One of the ELT-3 binding sites in F+G is overlapped with a noncoding DHS and footprint (dark green) (Gerstein et al. 2010). (B) Noncoding DHSs overlap two known elt-2 CRMs. Noncoding DHSs with TF footprints are detected upstream of elt-2 in two (CR I and CR III) of the three known elt-2 CRMs (purple boxes) (Wiesenfahrt et al. 2016). One footprint in the noncoding DHS overlapping CR III contains ELT-2 binding sites (TGATAA motifs; black).











