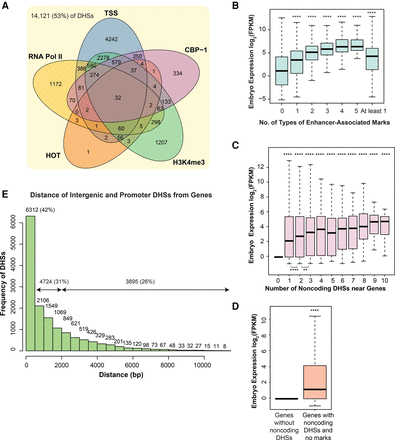
Presence of embryo noncoding DHSs near genes is associated with higher embryonic expression. (A) Half of embryo noncoding DHSs overlap with TSS, histone marks, CBP-1, and HOT regions. 47% of noncoding DHSs with marks of enhancer activity such as RNA Polymerase II (RNA Pol II; yellow), transcription start site (TSS; blue), CBP-1 (pink), H3K4me3 (green) observed in embryos, and modENCODE high occupancy TF regions (HOT; orange). TSS data are from Chen et al. (2013), and remaining data are from modENCODE (Gerstein et al. 2010). (B) Genes with noncoding DHSs harboring enhancer-associated marks are more highly expressed than those lacking any marks. Genes near embryo noncoding DHSs with any number of marks (at least one, two, three, four, and five type[s] of enhancer-associated mark) exhibit, on average, 8.9-fold higher levels of embryo expression (measured in log2 of FPKM) (data from Gerstein et al. 2010) compared with those with embryo noncoding DHSs lacking marks (P < 3 × 10−16). Genes with just one mark have 5.1-fold higher expression than genes without marks (P < 2 × 10−16). With each additional mark, median observed expression increases, up to three marks (5.1-fold higher expression compared with one mark, P < 3 × 10−14). No significant difference is observed between genes near noncoding DHSs with three, four, or five types of marks. (C) The presence of at least one embryo noncoding DHS near a gene is correlated with 4.5-fold higher embryo expression. The presence of at least one embryo noncoding DHS near a gene is associated with 4.5-fold higher embryo expression compared with genes without any DHSs (P < 3 × 10−16). Embryo expression (measured as log2 of FPKM) (data from Zhong et al. 2010) increases 54% from one to two embryo noncoding DHSs (P < 3 × 10−6) and 44% from two to three (P < 0.007). Further increases in DHS number are not correlated with increased expression. (D) Genes associated with embryo noncoding DHSs and lacking marks are still twice as highly expressed as genes without DHS. Genes with embryo noncoding DHSs lacking enhancer-associated marks (orange) show 2.3-fold higher embryo expression compared with genes lacking any DHSs (blue; P < 3 × 10−16). (E) Additional evidence for distant CRMs. Over half (56%) of intergenic and promoter DHSs are found within 1 kb of the nearest gene, and most (74%) are within 2 kb. However, a quarter (26%) of intergenic and promoter DHSs are >2 kb away and 10% are >4 kb away.











