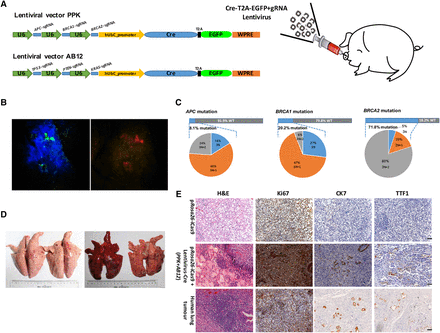
In vivo genome editing in the ear and lung tissues of Cre-dependent Cas9-expressing pigs. (A) Schematic of stereotactic delivery of lentiviruses AB12 and/or PPK into the ear and lung tissues of Cre-dependent Cas9-expressing piglets. (B) Florescence on ear tissues of Cre-dependent Cas9-expressing piglets infected with lentiviruses containing Cre, EGFP, and sgRNAs was directly observed using goggles. Left, EGFP fluorescence; right, tdTomato fluorescence. (C) Deep sequencing of sorted EGFP and tdTomato-positive cells show that all three sgRNAs could induce indel mutations near the predicted cleavage sites (8.10% of APC, 20.20% of BRCA1, and 71.80% of BRCA2), but not empty lentivirus infected cells. (D) Picture of the lungs from sacrificed Cre-dependent Cas9 expressing pigs infected (right) or uninfected (left) with lentivirus PPK and AB12. (E) Representative lung H&E staining and immunohistochemistry images of Cre-dependent Cas9-expressing pig injected with lentivirus PPK and AB12 at 3 mo posttransduction; tumor cells were stained positive for ki67 (an indicator of active cell cycle), CK7, and TTF1. Scale bar, 50 μm.











