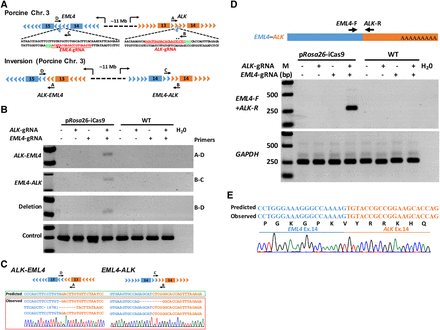
Induction of EML4–ALK rearrangements in pRosa26-iCas9 fibroblasts. (A) Schematic representation of porcine EML4–ALK rearrangements induced by CRISPR-Cas9. EML4-sgRNA and ALK-sgRNA (red) were designed to target the mutation sites of the porcine EML4 gene intron 14 and porcine ALK gene intron 13. PCR primers are indicated (primers A, B, C, and D). (B) PCRs were performed to analyze ALK–EML4 (primers A and D were used) and EML4–ALK rearrangements (primers B and C were used) and large fragment deletion (primers B and D were used). The fragment amplified by primers A and B was used as positive control (bottom panel). (C) The ALK–EML4 and EML4–ALK PCR products were subcloned into T-vector, and the Sanger sequencing results of five independent clones and a representative chromatogram are shown in the left and right panels, respectively. (D) Diagram of EML4–ALK mRNA fusion transcripts (upper panel). Agarose gel electrophoresis analysis suggested that the RT-PCR products of EML4–ALK mRNA fusion transcripts only exist in pRosa26-iCas9 fibroblasts infected with both EML4-sgRNA and ALK-sgRNA; GAPDH was used as positive control (bottom panel). (E) The Sanger sequencing results of RT-PCR products showing that the sequences of EML4–ALK mRNA fusion transcripts are identical with predicted sequences (bottom panel).











