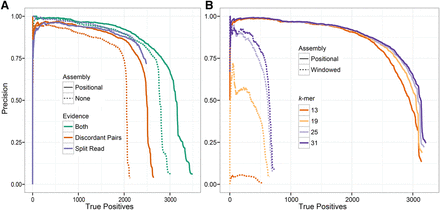
Performance of GRIDSS variant calling and assembly on NA12878 deletions events using long-read orthogonal validation data. Precision versus the number of true positives for different types of support (A) and for different k-mer sizes (B). Assembly of both split reads and read pairs improves both sensitivity and specificity to levels not achievable by either evidence source. Scoring only assembly-supported variants and varying the type of assembly and k-mer size demonstrates that robust small k-mer break-end assembly can be achieved with positional de Bruijn graph assembly but not windowed de Bruijn assembly.











