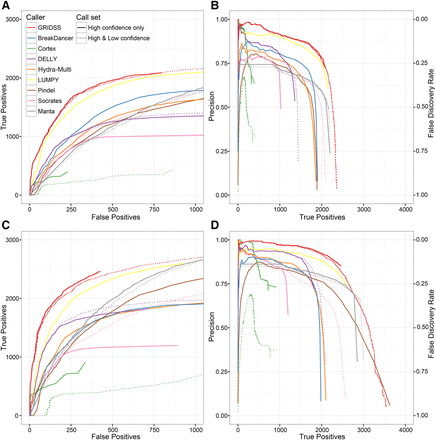
Performance of different SV callers on deletion detection in NA12878 at 50× coverage. Multiple variant calls were compared to both the Mills et al. (2011) validation call set (A,B) and PacBio/Illumina Tru-Seq Synthetic Long-Read (Moleculo) orthogonal validation data (C,D). Plots show the number of true positives versus false positives (A,C) and the precision versus true positives (B,D). Long-read validation required three split, or seven spanning long reads supporting the call.











