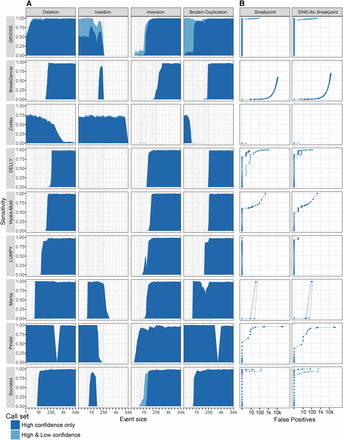
Variant caller performance on simulated heterozygous genomic rearrangements. Different classes of genomic rearrangement were randomly generated against human Chr 12 (hg19), and 60× coverage of 2×100-bp sequencing data was simulated. (A) The sensitivity of each method (rows) for each event type (columns) is plotted against event size. (B) Receiver operating characteristic (ROC) curves for all breakpoints (left) and breakpoints located in SINE/Alus (right).











