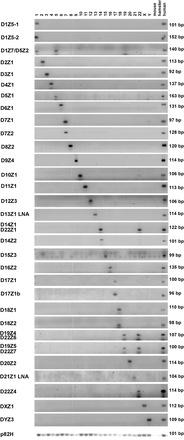
Detection of centromere α-repeat arrays in individual human chromosomes. Representative α-repeat arrays in each human chromosome (y-axis) were detected and the number of repeats quantitated by qPCR using specific primers. Gel electrophoresis of qPCR products amplified from DNA of human/rodent hybrid cells, with each hybrid cell containing only one human chromosome (displayed on the x-axis). DNA from rodent parental mouse or hamster cells is included to control for cross-species hybridization of repeats along with human DNA isolated from peripheral blood lymphocytes that served as a positive control. Water was used as an additional negative control. Using the primers and qPCR conditions described (Supplemental Tables 1, 2), specific centromeric α-repeat arrays were identified for each human chromosome (i.e., D2Z1, D3Z1, D4Z1…). Certain α-repeat arrays were found in two or more chromosomes (i.e., D1Z7/D5Z2 in Chromosomes 1 and 5, D14Z1/D22Z1 in Chromosomes 14 and 22, and D19Z4/D22Z6 and D19Z5/D22Z7 in Chromosomes 19 and 21). Primers specific for the ubiquitous α-repeat p82H amplified centromeres from all human chromosomes. Assays for the D13Z1 and D21Z1 arrays in this figure use LNA primers as shown in Supplemental Figures S4 and S5. The data shown in this figure are a composite from experiments run over time and demonstrate the results obtained once conditions for each chromosome had been optimized.











