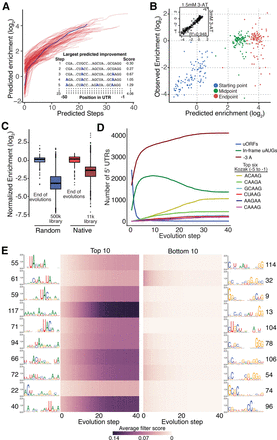
Model-guided optimization of 5000 random sequences. (A) Using our convolutional neural network, we iteratively predicted the optimal single nucleotide change in 100 random 5′ UTR sequences until no additional increase in enrichment was predicted. An example of these changes can be seen in the inset. (B) The start, midpoint, and endpoints from evolutions in A were tested experimentally. The predicted and observed enrichments are plotted. (C) Experimental data from endpoints of the optimized 5′ UTR sequences derived from both the random and native sets of sequence are compared to the enrichment distribution from the original random and native libraries. (D) Five thousand sequences from our random library were evolved over 40 steps and assayed for enrichment and depletion of common nucleotide features. (E) Analysis of the enrichment (left) and depletion (right) of motifs identified from the first convolutional layer of our model—the same as described in Figure 2.











