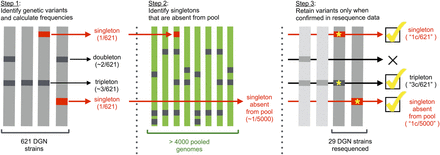
Pipeline for identification and validation of rare polymorphisms. Step 1 data set is from the Drosophila Genome Nexus (DGN) (Lack et al. 2015) which represent predominantly monoallelic genomes (i.e., either haploid or inbred) from 35 populations across three continents that were sequenced to high depth and underwent the same iterative mapping pipeline before variant calling. Step 2 data set consists of pooled sequencing data generated by our and collaborating labs which collectively represent >4000 genomes from the eastern US and Europe. Step 3 data set is resequence data made available by the Drosophila Genetic Reference Panel (DGRP) (Mackay et al. 2012) and DPGP1 (http://www.dpgp.org/1K_50genomes.html#Reference_Release_1.0; SRA accession number PRJNA3009) projects, which used Roche454 and Illumina technology (respectively) to independently resequence 29 of the strains present in the DGN.











