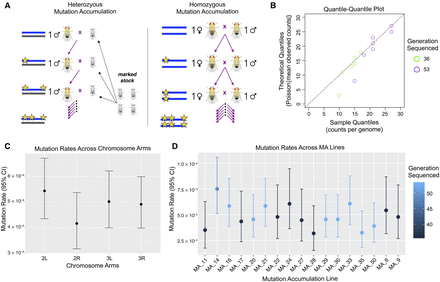
A summary of the experimental design and results for the single base pair mutation rate in this study. (A) Diagram depicting the general crossing schemes used in heterozygous (left) and homozygous (right) mutation accumulation, where this study used the heterozygous design. (B) QQ plot of the quantiles of the mutation counts on each chromosome arm of each strain, plotted against the quantiles of a Poisson distribution with mean taken from the mean counts in the MA experiment, where color indicates the generation sequenced (green = generation 36, purple = generation 53). (C) Mutation rates estimated for each chromosomal arm (Pearson's χ2 test of independence, χ2 = 2.55, df = 3, P-value = 0.47). (D) Mutation rates estimated for each strain, where color indicates the generation sequenced (Pearson's χ2 test of independence, χ2 = 7.99, df = 14, P-value = 0.89).











