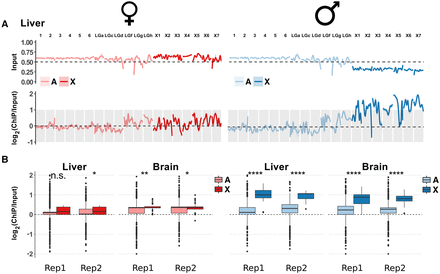
H4K16ac enrichment on the X Chromosome of Anolis males. (A) Genome-wide ChIP-seq profile along autosomes (light colors) and X-linked contigs (dark colors) for Anolis liver. Upper plots show data for the input (i.e., DNA prior to immunoprecipitation), which illustrate that the male X has only approximately half the coverage compared to autosomes and female X, consistent with the hemizygous state of male Anolis sex chromosomes. Lower plots display the input-normalized ChIP-seq signals and show the H4K16ac enrichment on the male X. All chromosomes are scaled to the same length; pink and light blue shaded areas around profile lines correspond to 95% confidence intervals. See Supplemental Figure S13 for the full data set (both organs, all replicates). Contig names correspond to those defined in the original Anolis genome paper (Alfoldi et al. 2011). (LG) Linkage group, (X1) LGb, (X2) GL343282.1, (X3) GL343338.1, (X4) GL343364.1, (X5) GL343417.1, (X6) GL343423.1, (X7) GL343550.1. (B) Comparison of H4K16ac distributions between autosomes and X Chromosomes along genic regions. Significance symbols are defined as in Figure 5. (Rep1) biological replicate 1; (Rep2) biological replicate 2. Comparisons along the whole genome and specifically for intergenic regions are reported in Supplemental Figure S12, A and B.











