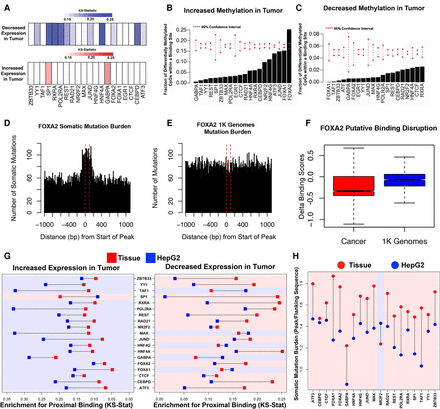
Primary tissue-derived DAP occupancy complements existing cell line data in characterizing liver cancer–associated genomic changes. (A) Color bars indicating the KS-test statistics for enrichment for binding proximal to the TSS of genes with significantly decreased (blue) or increased (red) expression in tumor tissue compared to adjacent normal tissue for each DAP. (B,C) Percentage of probes with significantly increased or decreased methylation in tumor compared to adjacent normal tissue overlapping a binding site of each DAP. Red dashed lines indicate 95% confidence intervals based on random sampling of an equivalent number of null probes. (D,E) Bars representing the number of somatic mutations (D) or matched 1000 Genome Project (E) mutations observed at contiguous 10-bp bins covering all FOXA2 peaks and flanking 1-kb regions. (F) Delta binding scores of all binding sites overlapping either somatic mutations found in cancer (red) or natural variation measured in 1000 Genomes (blue). (G) KS-test statistics for binding proximal to the TSS of genes with significantly increased or decreased expression in tumor tissue compared to adjacent normal tissue in HepG2 cells (blue) and adult liver tissue (red). (H) Mean somatic mutation burden of DAP binding sites over flanking regions in HepG2 cells (blue) and adult/male tissue (red). Regions are shaded according to whether HepG2- or tissue-derived peaks display greater enrichment.











