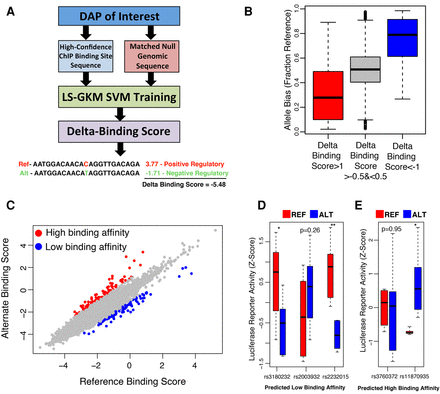
SNPs capable of disrupting regulatory activity can be identified from liver-derived data. (A) Diagram representing the pipeline for generating SVMs capable of distinguishing DAP binding sites from matched null regions and scoring the predicted impact of all possible mutations on each DAP. (B) Box plots of CTCF binding site overlapping, heterozygous SNPs predicted to be in the top ∼1% for decreasing binding affinity (red), to be in the top ∼1% for increasing binding affinity (blue), and to have no significant impact on binding affinity (gray). The y-axis indicates the fraction of ChIP-seq reads mapping to the reference allele. (C) SVM scores for reference and alternate GTEx liver eQTL SNP alleles for CTCF. Red dots indicate SNPs that hold a positive delta binding score in the top 0.1 percentile. Blue dots indicate SNPs that hold a negative delta-binding score that falls in the bottom 0.1 percentile of all scores. (D,E) Box plots representing the luciferase activity of reference (red) and alternate (blue) sequence in eQTL SNPs predicted to inhibit DAP binding (D) and induce TF binding (E). (*) Two-tailed t-test P < 0.05; (**) P < 0.005.











