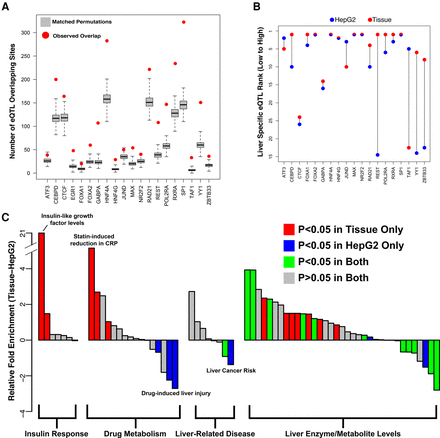
DAP occupancy is enriched for relevant trait- and expression-associated sequence variation. (A) Red dots indicate the number of eQTLs falling within a DAP binding site relative to the gray box plots, which represent 1000 randomly sampled null SNPs matched for distance to TSS and minor allele frequency. (B) Relative rank of liver-specific eQTL compared to all GTEx tissue specific eQTLs in DAP binding sites assayed in HepG2 cells (blue) and adult liver tissue (red). (C) Difference in enrichment (delta Fisher's exact test odds ratio) for SNPs associated with liver-related phenotypes between binding sites for 19 common DAPs assayed in HepG2 cells and liver tissue. GWAS terms represented by bars are provided in Supplemental Table 9A.











