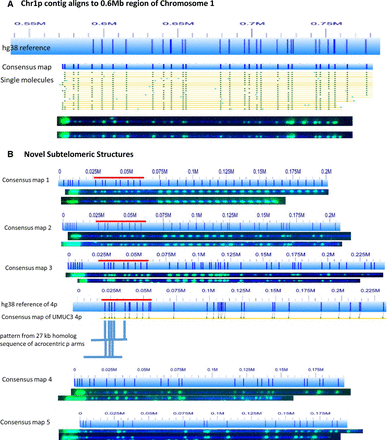
Telomere labeling identifies incorrectly represented and uncharacterized subtelomeres. (A) The 0.55- to 0.8-megabase region of the hg38 subtelomere reference sequence for 1p of UMUC3 is shown as a light blue bar; the dark blue vertical ticks within these bars indicate in silico Nt.BspQI nick-label sites. The hg38 reference sequence continuing toward the putative 1p telomere is not shown. The green lines on the single-molecule maps are Nt.BspQI (GCTCTTC) sites. Representative examples of raw images of several of these single molecules from two-color labeling experiments are shown. The single DNA molecules used to form the consensus map for Chromosome 1p each align with the 0.6-Mb region of the hg38 reference, and all contain intense telomere end labels. (B) Five telomere-containing consensus maps that could not be aligned with the hg38 reference sequence. Red bar designates an SRE located on 4p, 4q, and several acrocentric short-arm subtelomeres (Youngman et al. 1992).











