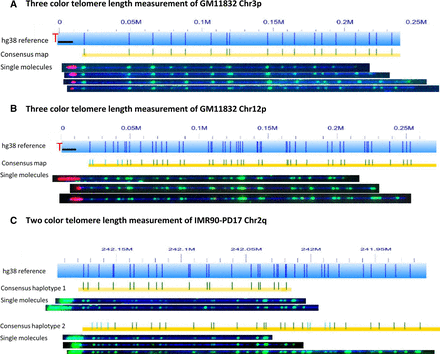
Colabeling telomere (TTAGGG)n tracts using gRNA-directed CRISPR/Cas9 nickase and subtelomeres using global motif-dependent nickase Nt.BspQI. The hg38 subtelomere reference sequences are shown as light blue bars; the dark blue vertical ticks within these bars indicate in silico Nt.BspQI nick-label sites. The black line segments designate telomere-adjacent gaps in the hg38 reference sequence. Individual single-molecule maps were de novo assembled into the consensus maps (yellow lines), which were then aligned with the hg38 reference (light blue bars). The green lines on the consensus maps are Nt.BspQI (GCTCTTC) sites that align to the corresponding reference site. Those that do not align with the reference are designated with light blue lines. Representative examples of raw images of single molecules that were used to create the consensus maps are shown. (A,B) Three-color consensus maps and single molecules for Chromosomes 3p and 12p of sample GM11832. The DNA backbone is blue, labeled subtelomeric Nt.BspQI sites are green, and the telomere (TTAGGG)n tracts are red (C) Two-color consensus maps and representative images of single molecules of both haplotypes for Chromosome 2q of sample IMR90-PD17. The DNA backbone is blue, and both the labeled subtelomeric Nt.BspQI sites and the telomere (TTAGGG)n tracts are green.











