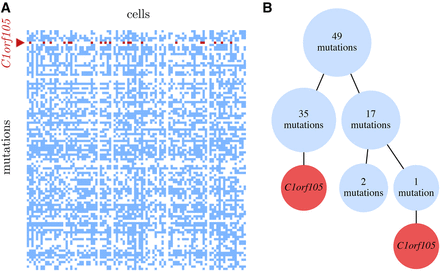
(A) The data matrix of the 105 mutations detected in the 96 single cells of patient 5 of the leukemia data set (Gawad et al. 2014). Unmutated positions are white, mutations are blue, and the recurrent mutation in C1orf105 is red. (B) The inferred mutational history under the finite sites model, when allowing a recurrence of the point mutation in C1orf105. The two occurrences appear at the ends of different lineages in the tree, separated in the two branches by 35 and 18 other mutations. The very large Bayes factor of 4.8 × 1015 shows that allowing the parallel mutation fits the data much better than enforcing the infinite sites assumption.











