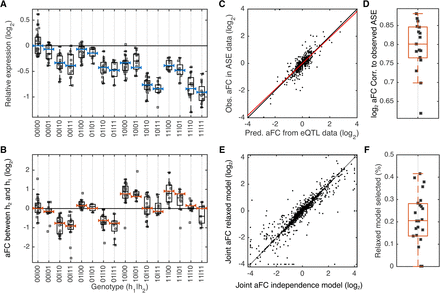
Joint analysis of aFCs for GTEx eGenes with two eQTLs. (A) An example of relative expression of eGene ZC3H3 and the model fits for different genotype groups of its two eQTLs (eVariant1: Chr 8: 144633728 A/G; eVariant2: Chr 8: 144556836 G/A) in GTEx adipose subcutaneous. The effect size of the first and the second eQTLs are −0.77 and −0.14 as measured by log2 aFC. Each dot represents observed expression in one individual, scaled relative to the expression at all-reference genotype. The blue bars show model fits from the two-eQTL model based on regulatory independence assumption. Reference and alternative alleles are denoted by 0 and 1, respectively, and haplotypes are separated by “|” sign (e.g., 10|11 corresponds to the cases that one haplotype carries alternative and reference alleles of eVariant1 and eVariant2, respectively, and the other haplotype carries the alternative allele of both eVariants). (B) Expression of the second haplotype relative to the first haplotype, observed in ASE data. The red bars show expected haplotype expression ratios based on the model in panel A, learned on the eQTL data. (C) aFC between two haplotypes as predicted from eQTL data compared with median aFC observed in ASE data for all eGenes with two eQTLs in adipose subcutaneous. Each dot represents one randomly selected genotype for one eGene. Red line indicates the robust linear fit (y = 0.9x + 0.002). (D) Predicted and observed median aFC for all eGenes with two eQTLs calculated from eQTL and ASE data, respectively, in each tissue with more than 200 eGenes with two eQTLs. (E) cis-Regulatory effect size associated with co-occurrence of the alternative alleles of the two eQTLs, as predicted under regulatory independence model or learned using the relaxed model. (F) Percentage of the two eQTLs that are not well described using the independent regulatory assumption across all tissues with more than 200 eGenes with two eQTLs.











