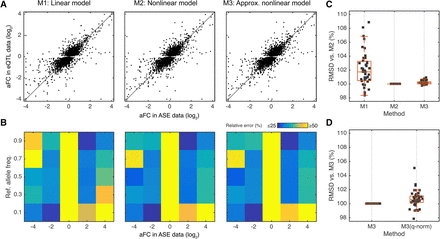
Comparison of the methods for estimating aFC using GTEx data. (A) aFC as estimated from ASE data versus estimates from eQTL data using linear model (M1), nonlinear model (M2), and the nonlinear model (M3) approximation for all top eQTLs in adipose subcutaneous. All three estimates are ∼75% correlated with estimates from ASE data. (B) Quality of the eQTL estimates as a function of allele frequency and the aFC estimate from allelic expression data, evaluated by average relative error between aFC from ASE data and from eQTL estimates. (C) Concordance between the estimates from allelic expression and eQTL data as evaluated by RMSD between the most accurate method, M2, and the other two methods. Each dot represents one tissue in GTEx. (D) Concordance between the estimates from ASE and eQTL data as evaluated by RMSD, comparing M3 to M3 applied after quantile normalization within each genotype group. Each dot represents one tissue in GTEx.











