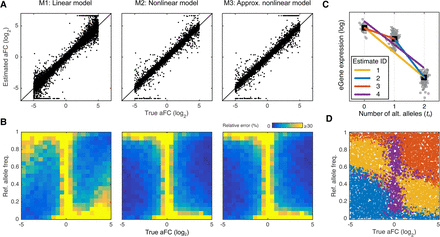
Comparison of the aFC estimation methods using simulated data. We simulated 10,000 eQTLs with noise (40% coefficient of variation), and uniformly selected log2 aFC (range: [−5,5]), and reference allele frequency (range: [0,1]). (A) True aFC used in simulation versus identified values using linear model (M1), nonlinear model (M2), and the nonlinear model approximation (M3). At this level of noise, M2 performed the best, with M1 and M3 having RMSDs of 164% and 110% of M2. (B) Quality of the effect size estimates as a function of allele frequency and the true effect size, evaluated by average error relative to the true log2 aFC. All three estimates, and particularly M1, deteriorate when the lower expressed allele is the minor allele. (C,D) Schematic representation of the nonlinear model approximation method (Box 3) based on four different candidate estimates (C), and the selected estimate with minimum residual variance for each simulated eQTL as a function of reference allele frequency and the true aFC (D).











