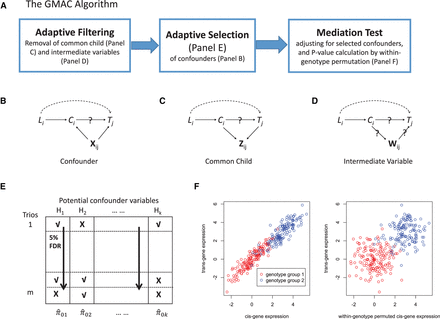
Graphical illustrations of GMAC and its main ideas. (A) A summary of the GMAC algorithm; (B) a mediation relationship among an eQTL, Li, its cis-gene transcript, Ci, and a trans-gene transcript, Tj, with confounders, Xij, allowing Li to affect Tj via a pathway independent of Ci; (C) a mediation trio where Ci and Tj have common child variable(s), Zij; (D) a mediation trio where Ci affects Tj through intermediate variable(s), Wij. (E) The adaptive confounder selection procedure: Based on the P-value matrix for the association of each potential confounder variable to at least one of the cis- or the trans-gene transcripts, we apply a stratified FDR approach by considering the P-values for each potential confounder (each column) as a stratum, with the significant ones indicated by a check mark (√). When conducting the mediation test for each trio, we only adjust for the significant confounding variables (the ones with √ in each row). (F) A mediation trio Li → Ci → Tj (left) and a trio under the null with both cis-linkage and trans-linkage but no mediation (right). Within-genotype permutation of the cis-gene expression levels maintains the cis- and trans-linkage (different mean levels) while breaking the potential correlation between the cis- and trans-expression levels within each genotype group. Note that Xij, Zij, Wij may vary by trios and are all subsets of H. We assume that either Xij or a combination of variables in Xij would capture the variation of the unmeasured confounder U in Figure 1.











