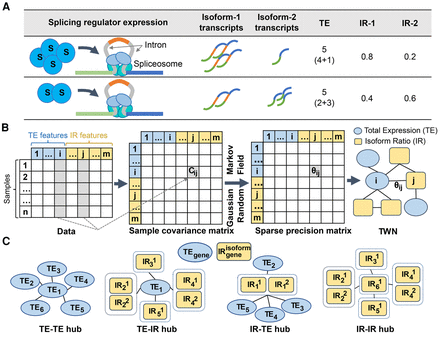
Transcriptome-Wide Network conceptual framework. (A) Schematic of the effect of a splicing regulator on inclusion of a cassette exon and resulting total expression and isoform ratios of the target gene. Splicing factor expression levels can affect splicing of target genes (Sveen et al. 2015). Higher expression of a splicing regulator S (first row) results in relatively more transcripts of isoform-1 and fewer of isoform-2. Total expression level is constant (5), but isoform ratios are different (0.4 and 0.6) as splicing factor S levels change (second row). (B) The (i,j)th cell of the sample covariance matrix contains covariance (Cij) between the ith and jth feature in data. We created a sparse precision matrix Θ (inverse covariance) from the sample covariance matrix using a graphical lasso to estimate the parameters of a Gaussian Markov random field. A nonzero value (Θij) in the precision matrix denotes an edge between the ith feature and jth feature in the network. (C) Edges in a TWN represent diverse relationships between total expression (TE) and isoform ratio (IR) nodes. Dotted rectangles group together isoform ratios for different isoforms of the same gene. Of particular focus are network “hub” nodes; in a TWN, there are four possible hub configurations depending on the node type of the central and neighboring nodes.











