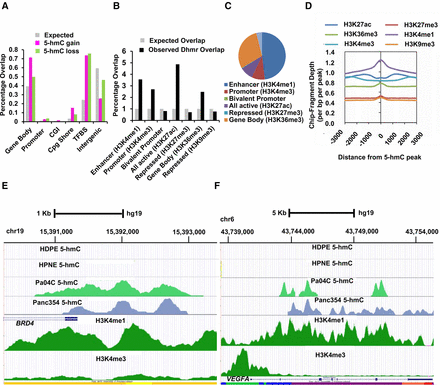
Pancreatic cancer cells acquire changes in 5-hmC at regulatory regions of the genome. (A) Common regions of 5-hmC gain and loss in pancreatic cancer occur significantly at gene bodies, promoters, and transcription factor binding sites. Gains/losses compared with expected genomic distribution (P-value <0.05). (B) Overlap of differentially hydroxymethylated regions in cancer (DHMR) with pancreatic histone marks shows enrichment at H3K4me1, H3K4me3, and other regulatory regions. (C) Distribution of differentially hydroxymethylated peaks shows most overlap with H3K4me1 regions. (D) Overlap of 5-hmC peaks with histone modifications shows spatial overlap with H3K4me1 regions. (E) BRD4 gene locus shows acquisition of 5-hmC peaks in pancreatic cancer cell line (Pa04C) and Xenograft (Panc354) around the promoter with strong overlap with H3K4me1 peaks. No 5-hmC peaks were seen in two healthy controls (HDPE, HPNE). (F) VEGFA gene locus shows acquisition of 5-hmC peaks in gene body with strong overlap with H3K4me1 peaks.











