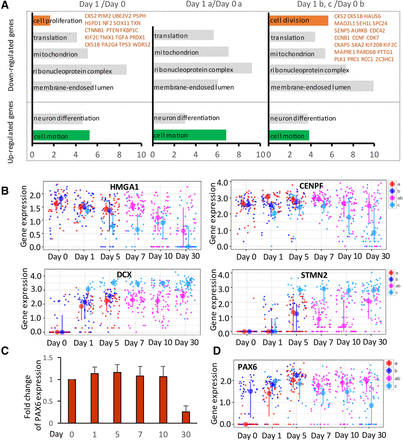
Subpopulation analysis reveals specific gene expression dynamics. (A) Gene ontologies associated with DEG comparing day 1 and day 0 (left), day 1 “a” and day 0 “a” subpopulation (middle), and day 1 “b” and “c” and day 0 “b” subpopulation (right) were generated by David analysis (Bioinformatics 6.7). The top GO terms are shown. The different GO terms are labeled with orange with representative genes. The x-axis indicates the −log (Benjamini P-value). (B) Dot plots show the representative cell cycle genes (HMGA1, CENPF) and neuron markers (DCX, STMN2) in different subpopulations as indicated for all six time points. The y-axis represents the gene expression level. Each small dot represents a cell, and the large dot indicates the medium expression of a given cell subpopulation. The error bar represents variation of the given gene in a given cell subpopulation. (C) Q-RT-PCR shows the expression level of PAX6 at six time points as shown, and the y-axis represents the fold change of PAX6 mRNA level. (D) The expression level of PAX6 in different cell subpopulations of different time points was shown in dot plots as in B.











