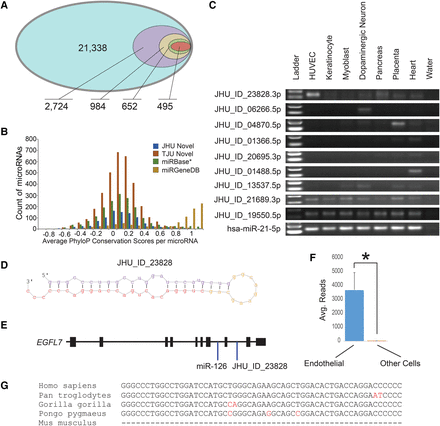
Some novel human microRNAs may still await characterization. (A) A nested Venn diagram was parsed down from 21,338 miRDeep2 identified novel microRNA loci to 2724 with ≥50 reads to 984 based on overlap with miRanalyzer-detected novel microRNAs to the 652 with both 5p and 3p mature microRNAs, and finally, with evidence of the RNA being Ago-bound, yielding 495 highest confidence novel microRNAs. (B) A histogram of PhyloP conservation scores averaged across the length of each mature microRNA. This collection of miRBase* v21 microRNAs has the miRGeneDB set removed. TJU novel microRNAs are from Londin et al. (2015). (C) Nine novel microRNAs were amplified that were predicted to be either cell-specific or ubiquitous. Most of these were lowly expressed. miR-21-5p was used as a control. (D) The predicted hairpin structure of novel microRNA JHU_ID_23828 is shown. (E) JHU_ID_23828 is located in the EGFL7 gene locus and shares a pri-miRNA with mir-126, an endothelial cell-enriched microRNA. (F) From the same sequencing batch, the average number of reads for JHU_ID_23828 among four endothelial cell types was 3673 and eight among 29 nonendothelial cell types ([*] P = 0.001, Mann-Whitney U test). (G) JHU_ID_23828 is present among primate species but is absent in lower mammals including Mus musculus.











