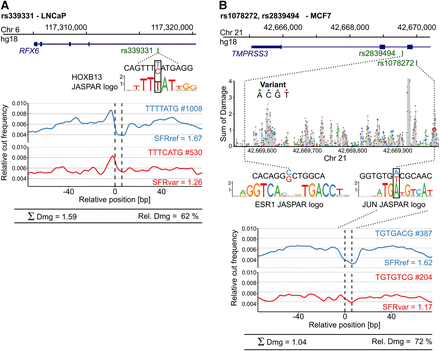
Assessing the footprint-damaging potential of functionally validated SNPs in nonerythroid tissues. Sasquatch-predicted damaging potential recapitulates functionally validated SNPs identified from GWAS studies in prostate and breast cancer. (A) SNP rs339331 T>C has been identified as significantly associated with prostate cancer risk and functionally validated to impair RFX6 expression by disrupting a HOXB13 binding site (Huang et al. 2014). Comparison of Sasquatch average profiles reflects the binding impairment and predicts a high damaging potential. (B) Two intronic SNPs (rs2839494 and rs1078272) found within an estrogen response element have been identified as significantly associated with survival in breast cancer patients and are functionally validated (Hsiung et al. 2014). Sasquatch predicts rs1078272 A>T to damage a potential JUN binding site, associated with a clear abolishment of a footprint in the average profile. In silico mutation identified rs1078272 to be located within a cluster of damaging variants. In contrast, rs2839494 C>G, proposed to directly alter ESR1 binding, is not detected. This is most likely because Sasquatch is limited in detecting large motifs spanning multiple noninformative bases, due to the use of short k-mers, which is also reflected in the lack of signal in the in silico mutation plot at that region. For the in silico mutation analysis, the T base at SNP rs107827 A>T retrieved from the hg18 reference genome has been changed from the minor to the major SNP allele.











