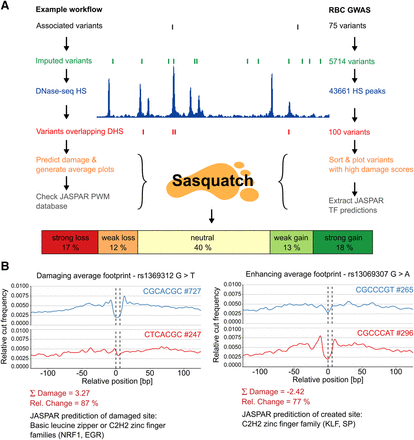
Prioritization of GWAS variants with Sasquatch. (A) The workflow for assessing and prioritizing GWAS variants according to their footprint-damaging potential is shown on the left. We employed Sasquatch using variants found to be significantly associated with red blood cell phenotypes (van der Harst et al. 2012). Starting from 75 GWAS-identified SNPs, variants were imputed and filtered for occurrence within genome-wide DHSs in primary erythroid tissue. For the 100 intersecting SNPs, damage scores were calculated and visualized using Sasquatch. (Note that variants not overlapping DHSs might still be interesting because of the potential introduction of novel TF-binding sites.) The top-ranked footprint-damaging (rs1369312 G>T) and enhancing variants (rs13069307 G>A) from this analysis are shown in B.











