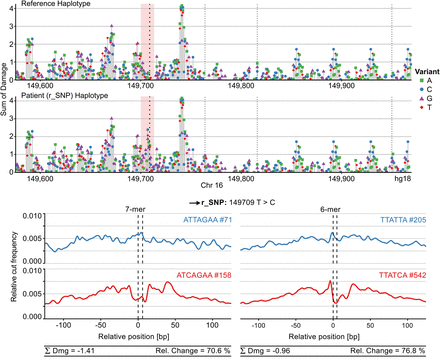
Characterizing a gain-of-function SNP. Sasquatch is capable of identifying a gain-of-function SNP associated with down-regulation of the HBA genes (De Gobbi et al. 2006) and characterizing its regulatory context using in silico mutation analysis. The black arrow indicates the position of the regulatory SNP (r_SNP). The in silico mutation plot identifies an introduction of a novel TF binding site (red panel) in the patient haplotype. Furthermore, the novel peak appears to lie adjacent to other potentially active binding sites, possibly contributing to the rSNP's regulatory context. Weaker dotted lines indicate known SNPs in the patient, neither of which appears to be associated with a strong change in footprint potential (see Fig. 5; Supplemental Fig. S1). The gain of a novel GATA footprint is captured both in 7-mer and 6-mer analysis, while 6-mers excel at capturing the characteristic shape of the GATA footprint.











