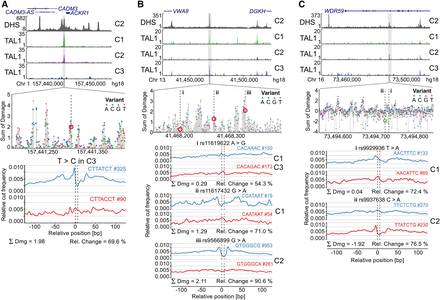
Prioritizing SNPs associated with differential TAL1 binding at single base-pair resolution. Several SNPs associated with differential TAL1 ChIP-seq peaks between three individuals (Lower et al. 2013) (C1, C2 and C3) were analyzed using Sasquatch. (A) A known T-to-C substitution at position −33 in the promoter region of the ACKR1 gene that abrogates its expression causing a form of the Duffy-negative genotype (de Carvalho and de Carvalho 2011). Sasquatch identifies a distinct depletion of a GATA1 footprint in the variant sequence. The variant lies within a cluster of damaging variants, strongly indicating a TF binding site, and is associated with abolishment of TAL1 binding in C3. (B) Three SNPs (rs11619622, rs11617432, and rs9566899) have been identified within a DHS on Chromosome 13 overlapping with TAL1 binding in C1 and C3 but not C2. Analysis of the rs11619622 A>G SNP present in C1 and C3 did not show striking differences between the reference and variant profile. The rs11617432 G>A SNP present only in C1 appears to disrupt a less common GATA binding motif, is predicted to have an intermediate damaging potential, but appears insufficient to abolish TAL1 binding. In contrast, the rs9566899 G>A SNP, found within a potential C2H2 zinc finger motif and present only in C2, shows the strongest predicted damage and appears sufficient to abrogate TAL1 binding in C2. (C) Sasquatch is able to predict the introduction of potential novel binding sites. Two SNPs (rs9929936 and rs9937638) have been found in an intragenic DHS associated with TAL1 binding only in C2. The rs9929936 T allele present in C1 potentially shows weak binding potential, but no alteration is caused by the variant. In contrast, the rs9937638 C>A SNP present in C2 shows the potential to introduce a novel GATA site. The sliding window approach identifies rs9937638 C>A to have a strong negative damaging potential in line with the observed TAL1 binding. Interestingly, the in silico mutation plot identifies nearby peaks of damage, potentially indicating bound motifs within the DHS that may support TAL1 binding.











