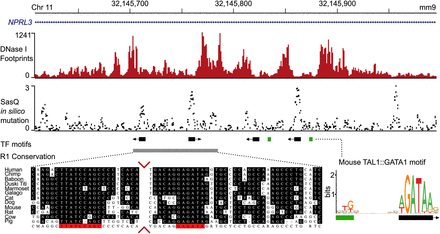
Overlay of DNase I footprints, Sasquatch in silico mutation predictions, and sequence conservation. The top panel shows deep DNase I footprints in mouse primary erythroid cells over the mouse alpha-globin locus R1 enhancer, which is present within an intron of NPRL3. Sasquatch's in silico mutation analysis using the same DNase-seq data reveals clusters of predicted high-damaging variants overlapping the footprints and known GATA1 binding motifs (middle panel). While sequence conservation analysis can identify the two leftmost binding sites, the in silico mutation analysis can also identify the murine-specific binding sites. Sequence conservation analysis adapted from Hughes et al. (2005). The conservation panel was trimmed to visualize both conserved sites (red arrows).











