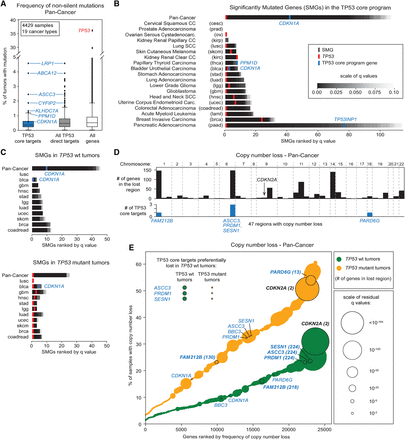
Select TP53 target genes are inactivated at low rates in human cancers expressing wild-type TP53. (A) Frequency of nonsilent mutations for TP53 core target genes, all direct TP53 target genes (early direct plus late targets with proximal binding), and all genes in the genome across 4429 tumor samples. (B) Significantly mutated genes (SMGs) in the TP53 core program identified by MutSigCV in pan-cancer and cancer-type–specific analyses. SMGs are ranked based first on q-values and then by mutation frequency. For the full list of SMGs, see Supplemental File S9. (C) SMG identification as in B dividing tumors based on their TP53 mutation status. Only tumor types with sufficient numbers in each group were analyzed. (D) Genome regions showing significant copy number losses identified with GISTIC 2.0 (q < 0.01) in a pan-cancer analysis. Lost TP53 core target genes are highlighted in blue. The complete list of significantly lost regions can be found in Supplemental File S10. (E) Cumulative distribution plots for copy number loss frequencies identified with GISTIC 2.0 in a pan-cancer analysis. The size of each point (gene) reflects the significance of the loss (q-value). TP53 wild-type (green) and mutant (orange) tumors were analyzed and plotted separately. Genes highlighted in bold show a significant copy number loss (GISTIC 2.0, q < 0.01). See also Supplemental Figure S6 and Supplemental Files S9, S10.











