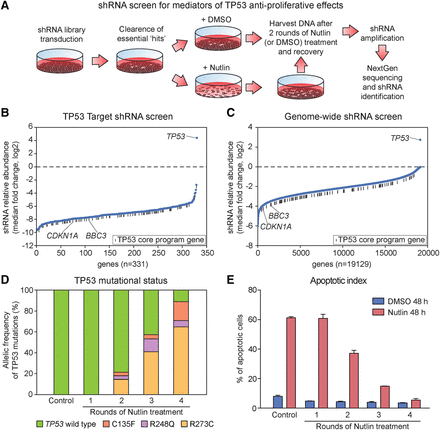
shRNA screens reveal that the anti-proliferative activity of TP53 is highly distributed among its target genes. (A) Schematic of experimental design. After library transduction, SJSA cells were propagated in culture to deplete cells carrying shRNAs targeting essential genes. SJSA cultures were then treated with Nutlin-3 or vehicle (DMSO) for 48 h, and surviving cells were then allowed to recover and propagate after drug removal. Two rounds of treatment and recovery were carried out before analysis of shRNA abundance. See Supplemental Methods for details. (B,C) Ranking of all genes targeted in each screen by median fold change (Nutlin-3/DMSO) of all shRNAs targeting each gene, from those showing strongest shRNA depletion to strongest shRNA enrichment. (D) Analysis of TP53 mutation status after multiple rounds of Nutlin-3 treatment and recovery as in A reveals the rapid appearance of inactivating mutations in the TP53 locus. (E) SJSA cells treated as in A become resistant to the apoptotic effects of Nutlin-3 after four rounds of treatment. After each round of recovery, cells were exposed to Nutlin-3 for 48 h and the fraction of apoptotic cells was measured by Annexin-V staining. See also Supplemental File S8.











