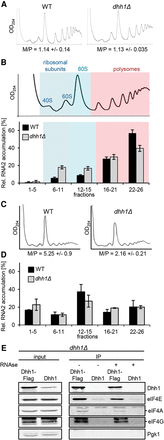
Depletion of Dhh1 shifts BMV RNA2 toward single ribosomal subunit fractions. (A) Global translation is unaffected in a strain expressing RNA2. UV absorbance profile at 254 nm of an extract from WT and dhh1Δ cells expressing RNA2 after sedimentation on a 10%–50% sucrose gradient. The monosome to polysome ratio (M/P ± SEM) is not significantly affected. (B) Sucrose density gradient analysis of BMV RNA2 in WT and dhh1Δ yeast. Below a representative UV absorbance profile is the distribution of normalized RNA2 levels in the specific fractions. Bars represent the average ±SEM from three independent experiments. Fractions were grouped into free (1–5), single ribosomal subunits (6–11), monosomes (12–15), light polysomes (16–21), and heavy polysomes (22–26). The total amount of RNA2 recovered over the gradient was set to 100%. (C) UV absorbance profile at 254 nm after 15-min glucose withdrawal of an extract from WT and dhh1Δ cells expressing RNA2. (D) Distribution of normalized RNA2 levels in the specific fractions after 15-min glucose withdrawal, as in B. (E) Dhh1 interacts with translation initiation factors. Western blot analysis of immunoprecipitation assays. Extracts were either treated (+) or not treated (−) with RNase A prior to the washing steps.











