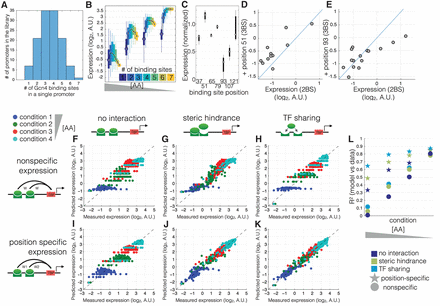
A model that incorporates TF sharing with specific position-expression can best explain expression across all amino acid concentrations. (A) The library consists of promoters with identical Gcn4 binding sites placed at one of seven locations in the promoter. (B) Shown are the measured expression levels (y-axis) as a function of binding site number (different colors) at four AA concentrations (different groups along the x-axis) for Gcn4. Each box contains data for all promoters with that number of binding sites and no other features (e.g., no nucleosome disfavoring sequences or binding sites for other TFs). The black line shows the median expression level for all promoters with that number of binding sites. (C) Shown is the expression for each promoter with a single Gcn4 binding site, normalized so that all conditions have the same mean expression. (D,E) Shown is the effect of adding a third binding site (at position 51 or position 93) to a promoter that already has two binding sites. The expression of the two binding site promoters (x-axis) is graphed against the three binding site promoters (y-axis). (F–L) Each point shows a single promoter measured at one of four conditions (blue, green, red, cyan in decreasing [AA] order) (x-axis) and the predicted expression levels (y-axis) of that promoter, for the six different models, fitted in cross-validation to the data shown in A, which are promoters with one to seven high-affinity Gcn4 binding sites (ATGACTCAT). R2 values were computed for absolute predicted expression on the test data. Each model includes either position-specific expression (a unique weight is associated with each unique binding site position) or nonspecific expression (all binding site positions share the same weight), and either no interaction, steric hindrance (a negative weight for multiple bound configurations), or TF sharing (the [TF] weight is divided by the number of sites). We note that the discretization of the y-axis in F–I is due to the fact that, in the absence of interactions and position-specific expression, all binding sites drive equal expression.











