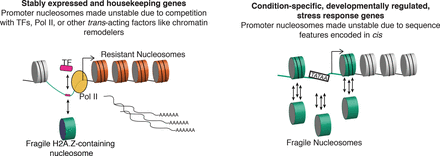
We propose a model whereby nucleosome fragility is determined by two distinct mechanisms: one that operates in cis at all genes and one that operates in trans at a subset of genes. (Left) Competition in trans with transcription factors and polymerase machinery destabilizes nucleosomes at the promoters of stably expressed genes. (Right) Condition-specific and developmentally regulated genes contain promoters with high levels of nucleosome fragility, determined primarily in cis by high AT content. (Green line) High AT content is sequence-encoded at all promoters but is highest at condition-specific genes; (orange cylinders) resistant nucleosomes found in the gene body of highly and stably expressed genes; (green cylinders) fragile nucleosomes compete (single arrow) with transcription factors and RNA Pol II at stably expressed genes. Fragile nucleosomes at condition-specific genes “treadmill” on the DNA (three arrows) due to destabilizing DNA elements like TATA-box motifs and high AT content.











