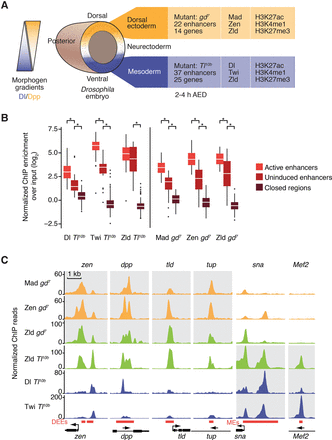
Dorsal-ventral (DV) transcription factors occupy uninduced enhancers at intermediate levels. (A) Overview of the model system of DV patterning in the Drosophila embryo, in which homogeneous cell fates can be obtained through mutants such as Tl10b and gd7, for which a large number of tissue-specific enhancers and their target genes are known. A summary of the analyzed ChIP-seq experiments of transcription factors and histone modifications is shown on the right. (AED) After egg deposition. (B) Boxplots of ChIP-seq enrichment over input for the DV transcription factors Dl, Twi, Mad, Zen, and Zld at known DV enhancers. The ChIP-seq experiments were performed in Tl10b or gd7 or both, dependent on which tissue the transcription factor is expressed in. Note that DV transcription factors occupy uninduced enhancers less than active enhancers but significantly more than closed regions, indicating that uninduced enhancers are accessible. Closed regions are 100 presumptive late enhancers that are inaccessible by DHS (Thomas et al. 2011) at early stages and are enriched for H3K27ac at later embryonic stages (see Methods). Active enhancers are mesoderm enhancers (MEs) in Tl10b embryos or dorsal ectoderm enhancers (DEEs) in gd7 embryos. Uninduced enhancers are MEs in gd7 embryos or DEEs in Tl10b embryos. Whiskers show 1.5 times the interquartile range, and outliers are shown as dots. Asterisk indicates P < 10−3 using the Wilcoxon rank-sum test. (C) ChIP-seq binding profiles of the transcription factors at four DEEs and two MEs (red boxes with target genes shown in black) illustrate higher binding at active enhancers (gray shading) but some degree of binding at uninduced enhancers (white background). The DEEs of zen, dpp, and tld are known to be repressed by Dl, while the snail (sna) ME is activated by Dl (Ip et al. 1992). ChIP-seq reads are normalized to reads per million.











