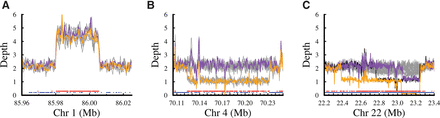
CNVs in this pedigree identified from nonplatinum variants. (A) Duplication on Chromosome 1 containing 242 Category 1 nonplatinum SNVs in a region of elevated read depth. Colored lines show the depths for the parents (NA12877 in purple and NA12878 in orange), and the gray lines show the depths for each of the children. Points along the bottom highlight the platinum variants (blue) and the nonplatinum variants (red). Note that there are 16 platinum variants in this duplication, because the presence of duplicated sequence can still produce genotypes that are consistent with those predicted by a diploid model (Supplemental Table S15). (B) Deletion on Chromosome 4 identified by 176 Category 2 nonplatinum SNVs that were consistent with the presence of a large hemizygous deletion. In addition to the large deletion, the depth supports the presence of several segmental duplications both overlapping and flanking the deletion. Lines and points are colored as in A. (C) Cell line or somatic deletions in multiple members of the pedigree on Chromosome 22 identified by 926 Category 4 nonplatinum SNVs. Lines are colored as in A, except for the addition of a black line that shows the depth for NA12893 (child). Although the other children (gray lines) do not appear to be deleted in any part of this region, the depths are highly variable, which may indicate somatic instability and mosaicism. The variability of this region within different cell line passages is evident when we compared these sequence data for NA12878 against corresponding data from the 1000 Genomes Project (1000 Genomes Project Consortium 2010) and NIST (Zook et al. 2014) (Supplemental Fig. S11).











