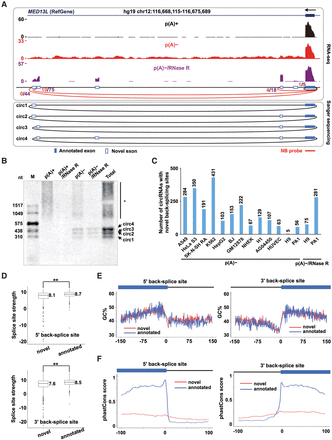
Unannotated exons produced from alternative back-splicing. (A) At least four novel exons (white bars) in the human MED13L locus were identified in PA1 p(A)− and/or p(A)−/RNase R RNA-seq data sets. The predicted circRNAs in the MED13L locus were indicated by red arc lines with raw back-splice junction reads from p(A)− (red) and/or p(A)−/RNase R (purple) RNA-seq data sets. Alternative back-splice sites were determined by both RNA-seq (top panel) and Sanger sequencing (bottom panel). Note that these new exons were barely detected in linear counterparts from parallel p(A)+ RNA-seq (the wiggle track in black). (B) Validation of multiple MED13L circRNAs with previously unannotated exons by Northern blot on native agarose gel. Note that the validation of these MED13L circRNAs with novel exons was consistent with RT-PCR (Supplemental Fig. S6B). (*) Linear RNA background. (C) Hundreds to thousands of circRNAs were identified with novel back-splice sites across different cell lines. (D) Splicing strength of novel back-splice sites is comparable to that of annotated back-splice sites. (**) P value < 0.01, Wilcoxon rank-sum test. (E) Novel (red) and annotated back-spliced exons (blue) have similar GC contents. (F) Novel back-spliced exons (red) are less conserved in sequences than are annotated back-spliced exons (blue).











