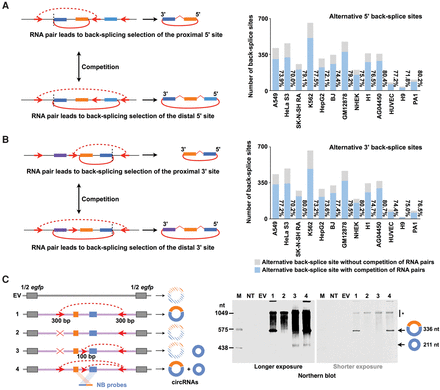
Competition of RNA pairs flanking proximal or distal back-splice sites leads to alternative back-splice site selection. (A,B) Potential RNA pairs (red dashed arc lines) produced by orientation-opposite complementary sequences (red arrows) flanking proximal (left top panels) or distal (left bottom panels) back-splice sites lead to alternative 5′ (A)/3′ (B) back-splice site selection, respectively (red arc lines). The competition of RNA pairs flanking proximal or distal back-splice sites leads to alternative back-splice site selection. More than 70% of the highly expressed circRNAs (RPM ≥ 0.1) with alternative back-splice site selection contain potential paired complementary sequences flanking both proximal and distal 5′/3′ back-splice sites (right panels). (C) Recapitulation of alternative back-splicing. (Left) A schematic drawing of egfp expression vectors with engineered complementary sequences for POLR2A circular RNA recapitulation. Half egfp sequences from the expression vector backbone are indicated as gray bars. POLR2A exonic and intronic sequences are indicated as colored bars and light purple lines, respectively. Nonrepetitive complementary sequences (red arrows) were inserted into multiple POLR2A intronic regions to form different RNA pairs (red dashed arc lines). Northern blot (NB) probes are indicated as colored bars. (Right) Validation of alternatively back-spliced POLR2A circRNAs by Northern blot on denaturing PAGE gel. Note that only partial complementary sequence (∼100 bp) was inserted into the middle intron for smaller POLR2A circRNA. (*) Linear RNA background.











