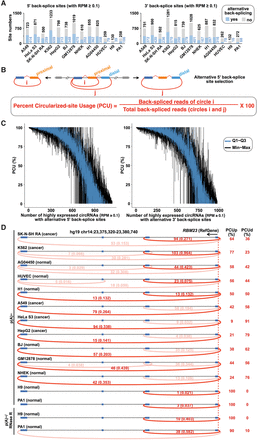
The diverse landscape of alternative back-splicing. (A) Approximately 12%–57% of back-splice sites are alternatively selected among high-confidence expressed circRNAs with RPM (mapped back-splice junction Reads Per Million mapped reads) ≥ 0.1. (B) A schematic diagram of alternative 5′ back-splicing and its quantification (top panel). The use of proximal and distal 5′ back-splice sites can be quantitated by the Percent Circularized-site Usage (PCU, bottom panel) with detected back-splice junction reads (i and j, respectively). (C) Diverse usage of alternative 5′ (left) and 3′ (right) back-splice sites among different cell lines. Each blue vertical line denotes PCU variation for one circRNA from the first quartile (Q1) to the third quartile (Q3) across cell lines, and each black vertical line denotes PCU variation from the minimum to the maximum. Note that only highly expressed circRNAs with RPM ≥ 0.1 in at least three cell lines were used for this analysis. (D) Visualization of alternative 5′ back-splice site usage in circRNAs produced from the RBM23 locus across different cell lines. Predicted circRNAs in the RBM23 locus were indicated by red arc lines with raw back-splice junction reads and normalized RPMs (numbers above each arc line, left panel). The use of proximal alternative 5′ back-splice site (PCUd) or distal alternative 5′ back-splice site (PCUp) was calculated accordingly (right panel).











