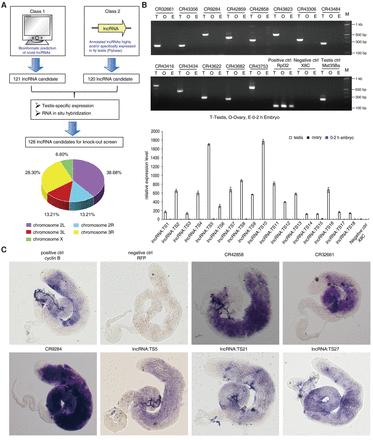
Systematic identification and validation of Drosophila lncRNAs involved in spermatogenesis. (A) Flowchart of identification and selection of testis-specific lncRNAs for the knockout study. Novel lncRNA prediction using bioinformatics and analysis of annotated lncRNAs in FlyBase were combined to build the lncRNA starting pool. Then, through a testis-specific expression screen and RNA in situ hybridization, 128 testis-specific lncRNA candidates were selected for targeted knockout. These lncRNAs were located on three different chromosomes, including the left and right arms of Chromosome 2, the left and right arms of Chromosome 3, and Chromosome X. (B) Testis-specific expression screen of predicted lncRNAs and annotated lncRNAs by quantitative RT-PCR and semiquantitative RT-PCR, respectively. RpL32 was used as an internal control. Values represent means ± SEM for three biological replicates. Mst35Ba was used as a testis-specific control. X8C (a Chromosome X-linked intergenic region that has been determined to be silent for transcription) was used as a negative control to rule out contamination of RNA by genomic DNA. (C) Expression of selected lncRNAs in Drosophila testis, analyzed by whole-mount in situ hybridization. Cyclin B RNA was used as a positive control, and RFP RNA in nontransgenic testis (w1118) was used as a negative control.











