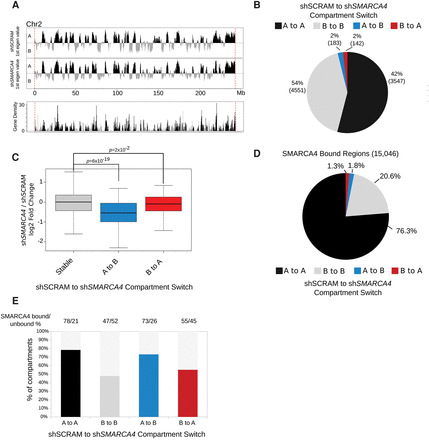
(A) Compartment profiles (the first principal components) of shSCRAM and shSMARCA4 data for Chr 2. The A-type (open) compartments are shown in black, and the B-type (closed) compartments are shown in gray. The same color scheme was used for the gene density plot for Chr 2 in the lower panel. (B) Pie chart showing the genomic compartment changes between shSCRAM and shSMARCA4 data sets. “A” and “B” denote the open and closed compartments, respectively. “A to A” represents compartments that are open in both cell lines; “B to B” represents compartments that are closed in both cell lines; “A to B” denotes compartments that are open in shSCRAM but closed in shSMARCA4; and “B to A” denotes compartments that are closed in shSCRAM and open in shSMARCA4. (C) shSMARCA4/shSCRAM log2 fold change RNA-seq expression box plot of all the genes residing at regions for different compartmental switch categories. The compartments that are switched from A to B and from B to A show significantly decreased and increased expression levels, respectively. P-value: Wilcoxon rank-sum test. (D) Pie chart showing the compartment-switching profiles of SMARCA4-bound regions. (E) Bar graph showing the percentage of the compartment-switching regions that are bound by SMARCA4. The colored portions of the graph denote the SMARCA4-bound percentage of each compartment-switching category.











