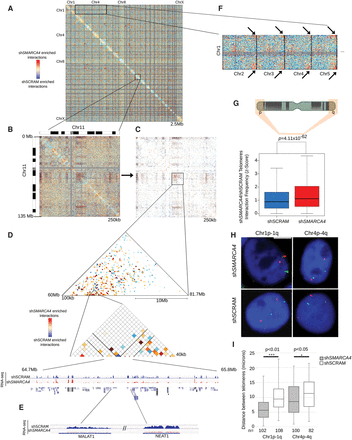
(A) Genome wide interaction heatmap at 2.5 Mb resolution showing the differences between interactions that are gained and lost upon SMARCA4 knockdown. The chromosomes are stacked from top-left to bottom-right in order (Chr 1, Chr 2, …, Chr 22, and Chr X). (B) A zoom-in of Chr 11 at 250-kb resolution showing all differential interactions. (C) The interactions that are altered with significance (see Methods). (D) A further zoom-in view of a genomic region on Chr 11 (Chr 11: 60000001–81750000) (top) showing the differential interactions where the MALAT1 and NEAT1 loci reside (Chr 11: 64750339–65807685). (E) RNA-seq tracks from shSMARCA4 and shSCRAM cells showing a reduction of expression in NEAT1 and MALAT1 lncRNA genes upon SMARCA4 knockdown. (F) A zoom-in of the inter-chromosomal interactions between Chr 1 and Chr 2 through Chr 5, with arrows indicating the enriched telomeric interactions in the shSMARCA4 cells. This pattern of subtelomeric interaction occurs throughout the genome. (G) Quantification of the interactions among subtelomeric ends for shSCRAM and shSMARCA4 Hi-C data sets. The subtelomeric ends show significantly (Student's t-test: P < 0.01 for Chr 1 and P < 0.05 for Chr 4) higher frequency of interactions in shSMARCA4 cells compared to control cells. (H) DNA-FISH images of shSMARCA4 and shSCRAM cells showing the intra-chromosomal telomeric interactions of Chr 1 and Chr 4. (I) Box plot showing the quantification of the telomere distances in shSMARCA4 and shSCRAM cells, quantified as described in the methods. P-value: Student's t-test.











