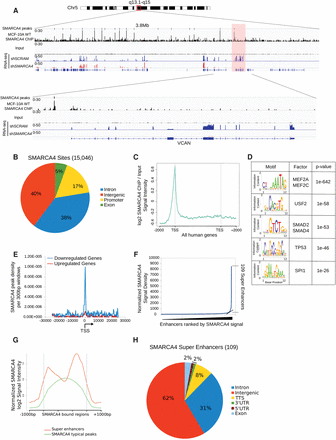
(A) Example of a ChIP-seq genome browser view of SMARCA4 binding and the input control, the shSCRAM and shSMARCA4 RNA-seq on Chr 5, and a zoom-in on the VCAN (Versican) gene, which is regulated by SMARCA4, in the lower panel. The y-axis represents the normalized tag densities relative to hg19 genomic coordinates. (B) Distribution of SMARCA4 ChIP-seq peak annotation for genic and intergenic regions. (C) Normalized SMARCA4 ChIP-seq signal intensity plot for all human UCSC genes ±2 kb. SMARCA4 binding is enriched at the promoter regions. (D) Top five sequence motifs associated with the SMARCA4 peaks. (E) SMARCA4 peak density within ±20 kb of the TSS of significantly down-regulated (blue), or up-regulated genes (red). (F) Distribution of SMARCA4 ChIP-seq signal across the MCF-10A enhancers. SMARCA4 binding is not uniformly distributed across the enhancers, as 109 super-enhancers display higher (log2 ∼1.5-fold) levels of SMARCA4 binding. (G) SMARCA4 signal is greater over super-enhancers (red) than typical enhancers (green). (H) Distribution of SMARCA4-bound super-enhancers in genic and intergenic regions.











