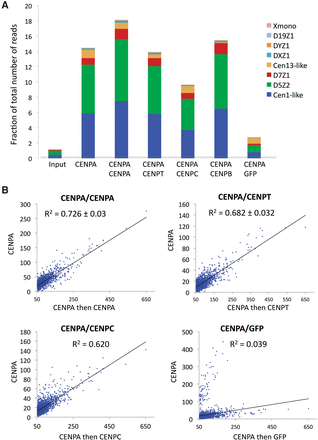
Sequential ChIP-seq data sets of CCAN components are enriched for centromeric α-satellites and are highly correlated. (A) Sequential ChIP-seq data sets show that abundant centromeric α-satellite arrays (D5Z2 and Cen1-like) are enriched for CENPA, CENPT, CENPC, and CENPB, whereas representative noncentromeric satellites (Xmono and D19Z1) are not. The reads were mapped to previously characterized α-satellite arrays (Hayden et al. 2013; Henikoff et al. 2015), and the number of reads mapped to an array were calculated (using Trim Galore!) as a fraction of the total number of mapped reads processed. (B) Close correspondence between sequential ChIP-seq data sets: CENPA/CENPA, CENPA/CENPC, and CENPA/CENPT. Scatter plots show the regression of the number of mapped reads for the CENPA/CENPA, CENPA/CENPC, CENPA/CENPT, and CENPA/GFP sequential ChIP data sets on the CENPA X-ChIP data set.











