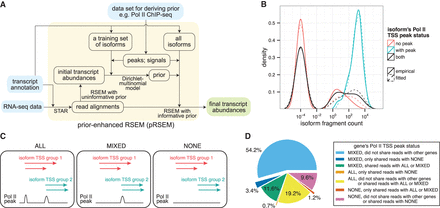
Pol II TSS peak data are informative for deriving a pRSEM prior. (A) pRSEM's workflow consists of inputs (light blue), intermediate steps (light orange), and output (green). (B) Empirical (solid lines) and fitted (dashed lines) distributions of fragment counts for isoforms in a pRSEM training set, stratified by Pol II TSS peak status. A small fractional count (10−4) was added to each isoform's fragment count so that isoforms with zero fragments could be depicted on a log scale. (C) Three types of gene Pol II TSS peak status. (D) Percentages of expressed genes classified by Pol II TSS peak status and multiread sharing. All fragment counts and percentages were calculated by RSEM on the K562 RNA-seq replicate one data set.











