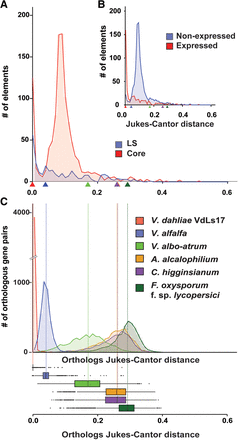
Dynamics of transposable elements in the genome of Verticillium dahliae strain JR2. (A) The divergence time of transposable elements identified in the genome of V. dahliae strain JR2 (Faino et al. 2015) was estimated using the Jukes-Cantor distance calculated between repeat copies and their consensus sequence. The distributions of divergence times between transposable elements located in the core genome (red) and in the LS regions (blue) differ. Estimations of speciation events in the evolutionary history of V. dahliae are indicated by triangles based on analyses in C. (B) The distributions of divergence times between expressed/active (log10[RPKM+1] >0) transposable elements (red) and nonexpressed (blue) transposable elements differ. Estimations of speciation events are indicated by triangles. (C) Speciation events are estimated by calculating the Jukes-Cantor distance for orthologous gene pairs based on genes from V. dahliae strains JR2 and their respective orthologs in the other genomes. Distributions and median divergence times between 1:1:1 orthologous pairs, displayed by box plots, were used to estimate relative speciation events.











