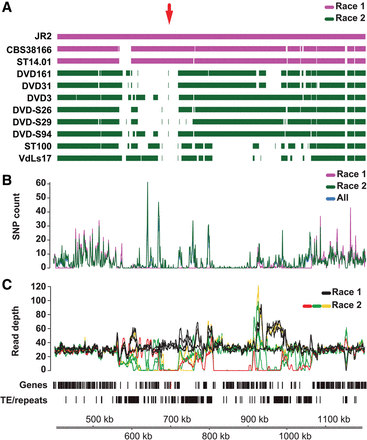
Details of the Ave1 locus in V. dahliae strain JR2. (A) Genome assemblies of race 1 and race 2 V. dahliae strains were aligned to the reference genome assembly of V. dahliae strain JR2. The red arrow indicates the location of the Ave1 gene. (B) Single nucleotide polymorphism (SNP) density (mean number of SNPs per 1 kb) over the Ave1 locus indicates depletion of SNPs in the Ave1 region when compared with neighboring regions. (C) A large genomic region on Chromosome 5 of V. dahliae strain JR2 containing the Ave1 gene is characterized by presence/absence polymorphisms between strains. Lines indicate the corrected average read depth (per 5-kb window, 500-bp slide) of paired-end reads derived from genomic sequencing of 11 V. dahliae strains. Different colors indicate distinct patterns of coverage across the Ave1 locus. Genes (Ave1 is marked in red) and transposable elements/repeats (excluding simple repeats) are indicated.











