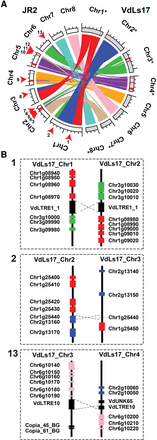
Extensive rearrangements in Verticillium dahliae genomes are mediated by repetitive elements. (A) Syntenic regions, indicated by ribbons, between chromosomes of the two highly similar V. dahliae strains JR2 (chromosomes displayed in white) and VdLs17 (chromosomes displayed in gray) reveal multiple synteny breakpoints caused by inter-chromosomal rearrangements, highlighted by red arrows for the JR2 genome. Red bars on the chromosomes indicate lineage-specific genomic regions (LS) that lack synteny in the other strain. To facilitate visibility, some chromosomes of V. dahliae strain VdLs17 have been reversed and complemented (indicated by asterisks). (B) Detailed view of the genomic regions surrounding selected synteny breakpoints. Rearrangements over short homologous regions such as repetitive elements (black boxes) or genes (colored boxes) resulted in inter-chromosomal rearrangements (translocations). V. dahliae strain VdLs17 genes were inferred by mapping of the V. dahliae strain JR2 genes to the genome assembly of V. dahliae strain VdLs17. Dashed gray lines indicate rearrangement sites. The numbers correspond to rearrangement numbers in A and Table 1.











